Breeding for parasite resistance in dairy cows

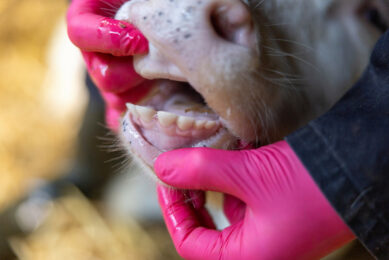
Gastrointestinal parasites can play havoc in the digestive tract of ruminants, causing a lack of appetite, diarrhoea and poor growth leading to economic losses for the farm. Selective breeding for parasite resistance in combination with other integrated control methods is considered an alternative means of parasite control.
Gastrointestinal parasites affect ruminants’ health, welfare and production and the quality of food resources. Despite all the traditional disease-controlling systems, parasites continue to impede the livestock industry. Approaches such as control of disease vectors and appropriate management methods reduce these negative effects. However, there are often constraints to the sustainability of such disease-control strategies, like environmental and food safety-related impacts of chemical treatments, the affordability and accessibility of treatments to poorer livestock producers, and the evolution of parasite resistance to the treatments.
Considering the economic impacts of health and welfare issues of parasites on the livestock industry, prevention of parasitic infections has more advantages than the cure. Ruminants’ resistance to gastrointestinal parasites is affected by different environmental and host factors, some metabolic diseases, host immune status, and pregnancy or lactation. Therefore, genetics of disease resistance involving immune and non-immune mechanisms is an approach to this issue.
Disease resistance
Disease resistance is defined as the inherent capacity of a previously unexposed animal to resist disease when challenged by pathogens or parasites. It is considered the host’s ability to moderate the pathogen or parasite lifecycle and resistance to the disease consequence of infection. Natural resistance is heritable from parent to offspring; therefore, increasing the overall level of genetic resistance of a population by using selective breeding programmes can improve animal health management systems.
Advantages of genetics
The benefits of incorporating genetic elements into disease management strategies include the permanence of genetic change once it is established, the consistency of the effect and the absence of the need for purchased inputs once the effect is established. Additionally, the effectiveness of other methods is prolonged as there is less pressure for the emergence of resistance.
There are also possible broad-spectrum effects, such as increasing resistance to more than one disease, the possibility of having less impact on the evolution of macroparasites such as helminths compared to other strategies such as chemotherapy or vaccination, and adding to the diversity of disease management strategies.
Management strategies
Applying different strategies to the genetic management of diseases depends on the nature of the problem and the resources available. These approaches include choosing the appropriate breed for the production environment, crossbreeding to introduce genes into well-adapted breeds to the required purposes, and selection for individuals with high levels of disease resistance.
There are breeding programmes focused on selecting commercial animals for enhanced resistance to some diseases such as parasite infections and some forms of mycotoxin poisoning. Finding genetic markers associated with resistance to infection potentially allows selection for increased resistance in the absence of infection.
Marker-assisted selection
Marker-assisted selection is a process in which a morphological, biochemical or DNA/RNA-based marker is used for indirect selection of a trait of interest such as disease resistance. The marker used for selection is associated at high frequency with the gene of interest due to genetic linkage. There are selectable markers, which eliminate certain genotypes from the population, and screenable markers, which cause certain genotypes to be readily identifiable, at which point the experimenter must score or evaluate the population and act to retain the preferred genotypes.
Depending on the trait of interest, marker-assisted selection may be cheaper and faster than conventional phenotypic assays. Multiple markers can be evaluated using the same DNA sample; once DNA is extracted and purified, it may be used for multiple markers, for the same or different traits, thus reducing the time and cost per marker.
Identifying markers
Genetic markers can be identified by a range of molecular techniques such as microsatellites or single nucleotide polymorphism detection. Linked markers are molecular markers located very close to major genes of interest. There are several known phenotypic and genetic markers for resistance to gastrointestinal parasites in naturally infected ruminants that could potentially assist response to selection.
The major histocompatibility complex (MHC) involves a series of highly polymorphic genes that are responsible for the initiation of the immune response when an animal is challenged by pathogens or parasites. The MHC is divided into 3 regions: class 1, class 2 and class 3. In ruminants, there is a division of the class 2 region into 2 subregions: class 2a and class 2b.
Defining what trait to measure
Faecal egg count (FEC) is used as an indicator trait to determine resistance to gastrointestinal parasites. Originally developed for sheep, it can also be used for cattle, pigs and poultry as well as other species. The heritability of the FEC varies considerably, depending on both the parasite species and animal breed. Immune response evaluation is another indicator trait that can be used.
Selective breeding
Selective breeding to take advantage of within-breed variation in disease resistance is an important disease control strategy. Selection for parasite resistance alone can result in negative traits such as lower live-weight gains. Therefore, it is essential to apply appropriate selection policies and to understand the genetic architecture of underlying resistance to predict genetic risk or selective breeding.
Conclusion
Application of selective breeding for parasite resistance in combination with other integrated control methods is considered an alternative means of parasite control in the long run. However, disease resistance varies between species, between breeds and between individuals within breeds. Identification of the phenotype for disease resistance is challenging, because in a population containing both healthy and sick animals, not all healthy animals may be disease-resistant.
Join 13,000+ subscribers
Subscribe to our newsletter to stay updated about all the need-to-know content in the dairy sector, two times a week.